Tagged: Digdag
Digdag is a simple tool that helps you to build, run, schedule, and monitor complex pipelines of tasks. Its Simple, Open Source, Multi-cloud workflow engine.
In Digdag, workflows are typically defined in YAML files with a “.dig” extension. Developers usually write these workflows, which consist of tasks to be executed. However, tasks can also be added dynamically using the Digdag Python API or by downloading a “.dig” file from a remote HTTP server and incorporating it as a subtask. This approach is useful when a web service or app generates customized workflow files based on web app conditions, allowing the workflow logic to be managed externally. You can add webhooks to make it reactive.
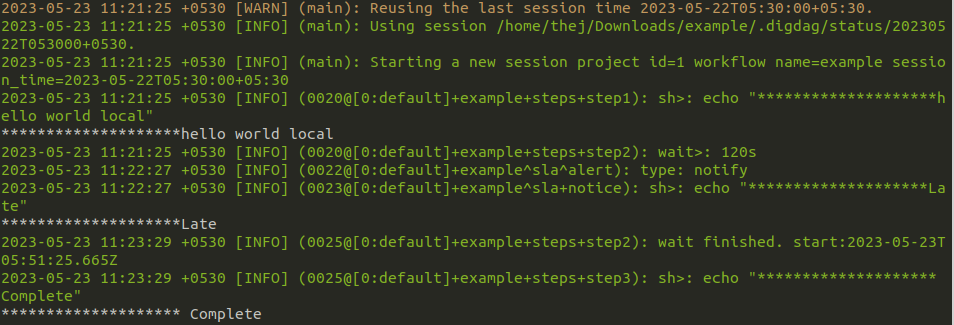
You can set up a failure alert task _error in Digdag to alert you when a workflow fails. But sometimes you want to get an alert even if the task runs successfully but takes more time than expected. For this, you can use sla feature.
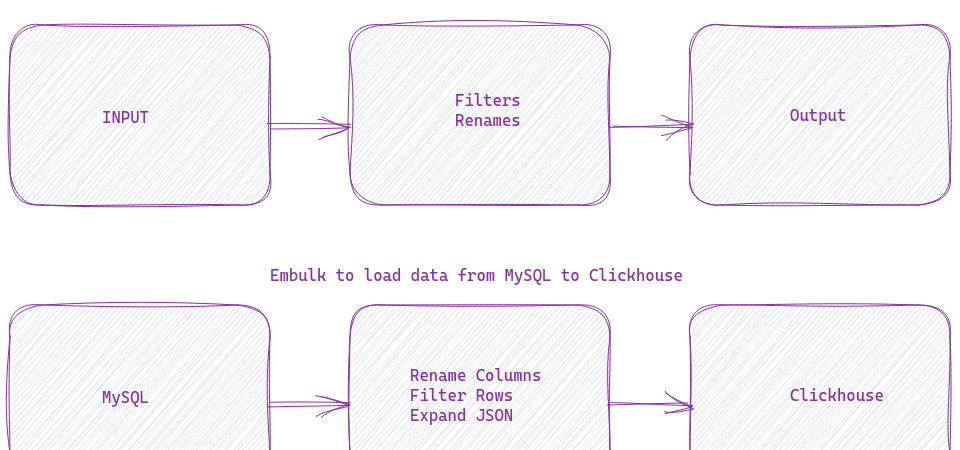
Embulk needs a YAML file configuration for each data load. It’s a simple format, very human-readable. But there are cases where I want the YAML files to generate dynamically. Embulk does support an experimental feature that involves liquid templates. But my team is well versed in Python and Jinja2. Hence that is what we use.
Embulk is a bulk data loader. It helps transfer data between different types of databases, storages, file formats, cloud services, etc. It’s like a Unix tool. It’s simple, robust, and works well with other tools.
I often try to use Docker with my Digdag workflows. That way, we can keep the dependencies clean and separate for each DAG. I have blogged about it in detail before. Now instead of Docker, I can use Podman, by just making one small change.
There is a good chance that you want to allow multiple developers to run their dags/tasks on your Digdag server. It’s painful to maintain the packages/dependencies they need to run their tasks. It specially becomes difficult if they conflict with each other. One way to solve it, is by having some kind of...